RESAMPLING METHODS, PART I – JACKKNIFE METHODS
1. Validation Set Approach
> attach(Auto);
n = length(mpg);
> Z = sample(
n, n/2 ) # Random subsample of size n/2
# Works similarly to generating a Bernoulli variable
> reg.fit
= lm( mpg ~ weight + horsepower + acceleration, subset=Z ) # Fit using training data
> mpg_predicted
= predict( reg.fit, Auto )
#
Use this model to predict the testing data [-Z]
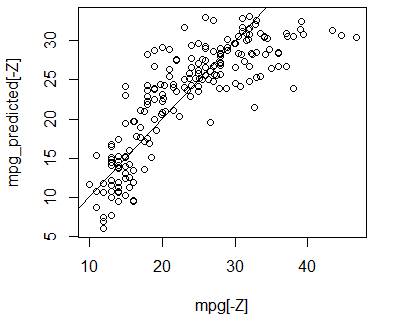
> plot(mpg[-Z],mpg_predicted[-Z]) # We can see a nonlinear component
> abline(0,1) # Compare with the line y=x.
> mean(
(mpg - mpg_predicted) [-Z]^2 ) #
Estimate the mean-squared error MSE
[1] 20.46885
2. Jackknife (Leave-One-Out Cross-Validation,
LOOCV)
> install.packages(“boot”)
> library(boot)
> glm.fit
= glm(mpg
~ weight + horsepower + acceleration) # Default “family” is Normal, so this GLM
model
> cv.error
= cv.glm(
Auto, glm.fit )
# is the same as LM, standard linear
regression.
> names(cv.error)
# This cross-validation tool has several
outputs
[1] "call"
"K"
"delta" "seed" # We are interested in “delta”
> error$delta
[1] 18.25595 18.25542
# Delta consists of 2
numbers – estimated prediction error and its version adjusted for the lost
sample size due to cross-validation.
# Example – what power of “horsepower” is
optimal for this prediction?
> cv.error
= rep(0,10) # Initiate a vector of estimated errors
> for
(p in 1:10){ # Fit polynomial regression models
+ glm.fit
= glm(
mpg ~ weight + poly(horsepower,p) + acceleration ) #
with power p=1..10
+ cv.error[p]
= cv.glm(
Auto, glm.fit )$delta[1] } # Save prediction errors
> cv.error # Look at the results
[1] 18.25595 15.90163 15.72995 15.86879 15.74517
15.74989 15.70073 15.79314 15.95933 16.38301
# Although p=7 yields
the lowest estimated prediction error, after p=2, the improvement is very
little.
> plot(cv.error)
> lines(cv.error)
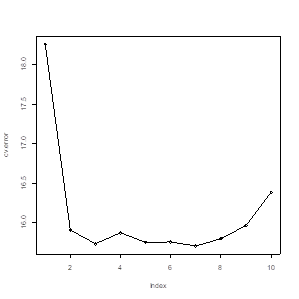
# Package “bootstrap” has a jackknife tool.
For example, here is a bias reduction for a sample variance.
# We’ll start with the
biased version of a sample variance (it’s MLE, under Normal distribution)
> variance.fn = function(X){ return( mean(
(X-mean(X))^2 ) ) }
> variance.fn(mpg)
[1] 60.76274
# Use jackknife to correct its bias.
> install.packages(“bootstrap”)
> library(bootstrap)
> variance.JK
= jackknife( mpg, variance.fn
)
# This gives us the
jackknife estimates of standard error and bias, as well as all estimates ![]() (-i)
(-i)
> variance.JK
$jack.se
[1] 3.741951
$jack.bias
[1] -0.1554034
$jack.values
[1] 60.84210 60.73524 60.84210 60.77598
60.81160 60.73524 60.68936 60.68936 60.68936 60.73524 60.73524 60.68936
60.73524 60.68936 60.91735 60.91278 60.84210 60.90280
[19] 60.88575 60.90142 60.91195 60.91735
60.91195 60.90142 60.90280 60.45457 60.45457 60.52096 60.38306 60.88575
60.86496 60.91195 60.86746 60.77598 60.81160 60.86746
[37] 60.84210 60.68936 60.68936 60.68936
60.68936 60.58222 60.63836 60.63836 60.84210 60.91278 60.86746 60.84210
60.91763 60.86496 60.80800 60.80800 60.77182 60.57584
[55] 60.88575 60.90142
60.91735 60.91195 60.91763 60.88770 60.90280 60.63836 60.68936 60.73524
60.68936 60.81160 60.52096 60.63836 60.58222 60.63836 60.86746 60.73524
[73] 60.63836 60.63836 60.68936 60.84210 60.91278
60.90280 60.90142 60.91278 60.86496 60.91763 60.86496 60.88575 60.63836
60.68936 60.63836 60.68936 60.73524 60.58222
[91] 60.63836 60.63836 60.68936 60.63836
60.58222 60.63836 60.84210 60.77598 60.84210 60.84210 60.91763 60.90142
60.52096 60.58222 60.63836 60.58222 60.84210 60.88770
[109] 60.90280 60.91278
60.84210 60.86746 60.90280 60.90142 60.73524 60.77598 60.83905 60.91735
60.88770 60.86746 60.73524 60.91735 60.88770 60.52096 60.88770 60.86746
[127] 60.73524 60.77182
60.90142 60.73052 60.91195 60.77598 60.77598 60.84210 60.77598 60.63836
60.68936 60.68936 60.68936 60.83905 60.90142 60.90142 60.77182 60.73052
[145] 60.86496 60.91735
60.90142 60.91735 60.90142 60.77182 60.86746 60.84210 60.73524 60.73524
60.77598 60.73524 60.77598 60.68936 60.81160 60.77598 60.73524 60.84210
[163] 60.90280 60.88770
60.63836 60.83905 60.91763 60.88770 60.91763 60.91735 60.91195 60.91735
60.84210 60.83905 60.86746 60.91763 60.91763 60.91278 60.91195 60.68409
[181] 60.86496 60.91195
60.91195 60.90142 60.88575 60.82749 60.77598 60.75625 60.71294 60.91278
60.91278 60.91735 60.91585 60.83905 60.91529 60.83905 60.68409 60.88770
[199] 60.84210 60.85542
60.82749 60.82416 60.73052 60.86496 60.89423 60.88770 60.63836 60.86746
60.86746 60.79444 60.79444 60.63836 60.63836 60.63836 60.75181 60.80800
[217] 60.51403 60.90732
60.65895 60.82749 60.81160 60.75625 60.73524 60.82749 60.89589 60.86746
60.85542 60.77598 60.75625 60.75625 60.77598 60.83905 60.91529 60.90142
[235] 60.90732 60.79055
60.65895 60.80800 60.79055 60.91278 60.90843 60.90843 59.92768 60.50757
60.69379 60.26550 60.50757 60.88590 60.87617 60.89113 60.87192 60.89589
[253] 60.89113 60.91113
60.89589 60.87617 60.89737 60.90019 60.85793 60.84486 60.87192 60.83349
60.84486 60.82749 60.80800 60.87600 60.88201 60.77567 60.90403 60.91799
[271] 60.91782 60.91761
60.89277 60.81160 60.90940 60.78352 60.75181 60.82416 60.90843 60.88406
60.91477 60.89113 60.89737 60.81160 60.83051 60.79444 60.84758 60.80827
[289] 60.75625 60.87192
60.85542 60.73488 60.62709 60.53311 60.87805 60.90835 60.91763 60.88201
60.91761 60.62160 60.60483 60.73919 60.42600 60.85521 60.84464 60.88930
[307] 60.65895 60.08238
60.36752 60.72611 60.43308 60.86496 60.89577 60.91627 60.86971 60.61606
60.81462 60.75997 60.44709 60.72165 59.54351 60.86727 60.14593 59.80304
[325] 59.89721 60.48787
60.80800 59.77073 60.64325 60.81462 60.69856 60.91798 60.57584 60.71256
60.88201 60.89263 60.90393 60.91813 60.80800 60.28981 60.29781 60.56989
[343] 60.71713 60.44709
60.39717 60.62709 60.59339 60.61047 60.81133 60.68409 60.64854 60.71256
60.68896 60.74766 60.86260 60.78322 60.90835 60.91668 60.91534 60.89263
[361] 60.89113 60.83051
60.86496 60.88575 60.63253 60.77182 60.83905 60.88575 60.91735 60.51403
60.44709 60.77182 60.37501 60.51403 60.51403 60.51403 60.63253 60.37501
[379] 60.73052 60.37501
60.91195 60.37501 60.90142 60.91278 60.73052 60.51403 60.88575 60.88575
59.83489 60.73052 60.86496 60.77182
# We can now calculate
the jackknife variance estimator by the jackknife formula
> variance.fn(mpg) - variance.JK$jack.bias
[1] 60.91814
# … and this is precisely the usual, unbiased
version of the sample variance.
> var(mpg)
[1] 60.91814
3. K-Fold Cross-Validation
# We can specify the number of folds K
within cv.glm (Omitted K is K=n by default, which is
LOOCV).
> cv.error
= rep(0,10)
> for
(p in 1:10){ glm.fit = glm(
mpg ~ weight + poly(horsepower,p) + acceleration )
+ cv.error[p]
= cv.glm(
Auto, glm.fit, K=30 )$delta[1]
}
> cv.error
[1] 18.22699 15.87030 15.73565 15.85911
15.80686 15.71585 16.09625 15.67797 15.82097 16.48196
> which.min(cv.error)
[1] 6
4. Cross-validation in classification problems.
Loss function.
# Command cv.glm, as
we used it above, calculates MSE, the mean squared error, and calls them
“delta”.
# In classification
problems, MSE can be used to measure the distance between the response variable
Y
# and the predicted
probability p. For this, Y has to be 0 or 1, and p
should be the probability of Y=1.
# However, the correct
classification rate and the error classification rate are more standard
measures
# of classification
accuracy. We can force cv.glm to return these
measures by introducing a suitable loss
# function. For
example, we’ll be predicting whether a student is
depressed or not.
# We define a loss function L(Y,p), a function of true response Y and predicted
probability p. The loss = 1 if
# the predicted response is different from the
actual response and 0 otherwise. Suppose the threshold is 0.5
> loss = function(Y,p){ return( mean( (Y==1 & p < 0.5) | (Y==0 & p
>= 0.5) ) ) }
> loss(1,0.3)
[1] 1
> loss(c(1,1),c(0.3,0.7))
[1] 0.5
# Now we attach the
Depression data, skip missing values, fit logistic regression model, and
estimate
# the error
classification rate by LOOCV.
> Depr
= read.csv(url("http://fs2.american.edu/~baron/627/R/depression_data.csv"))
> D = na.omit(Depr)
> attach(D)
> library(boot)
> lreg = glm(Diagnosis ~
Gender + Guardian_status + Cohesion_score,
family="binomial")
> cv
= cv.glm( D, lreg, loss )
> cv$delta[1]
[1] 0.1615721
# Is Guardian_status significant?
Let’s compare the error rates with and without
variable “Guardian_status”.
> lreg = glm(Diagnosis ~
Gender + Cohesion_score, family="binomial")
> cv.glm( D, lreg, loss )$delta[1]
[1] 0.1572052
# The error
classification rate is lower without the “Guardian_status”.